- Tidy Data
- Spreading and Gathering
- Separating and Uniting
- Missing Values
- Comparing the
fillarguments ofspread()andcomplete() - Nontidy Data
** This post is heavily based on R for Data Science. Please consider to buy that book if you find this post useful.**
tidyr, a member of the core tidyverse, will be used in this chapter.
Tidy Data
The tidy dataset, will be much easier to work with inside the tidyverse.
There are three interrelated rules which make a dataset tidy:
- Each variable must have its own column.
- Each observation must have its own row.
- Each value must have its own cell.
That interrelationship leads to an even simpler set of practical instructions:
- Put each dataset in a tibble.
- Put each variable in a column.
The two main advantages of tidy data:
- Due to having a consistent data structure (i.e., tidy data), it’s easier to learn the tools that work with it because they have an underlying uniformity.
- There’s a specific advantage to placing variables in columns because it allows R’s vectorized nature to shine. Most built-in R functions work with vectors of values. That makes transforming tidy data feel particularly natural.
All of the packages in the tidyverse are designed to work with tidy data. For example:
# Compute rate per 10,000
table1 %>%
mutate(rate = cases / population * 10000)## # A tibble: 6 x 5
## country year cases population rate
## <chr> <int> <int> <int> <dbl>
## 1 Afghanistan 1999 745 19987071 0.373
## 2 Afghanistan 2000 2666 20595360 1.29
## 3 Brazil 1999 37737 172006362 2.19
## 4 Brazil 2000 80488 174504898 4.61
## 5 China 1999 212258 1272915272 1.67
## 6 China 2000 213766 1280428583 1.67# Compute cases per year
table1 %>%
count(year, wt = cases)## # A tibble: 2 x 2
## year n
## <int> <int>
## 1 1999 250740
## 2 2000 296920# Visualize changes over time
library(ggplot2)
ggplot(table1, aes(year, cases)) +
geom_line(aes(group = country), color = "grey50") +
geom_point(aes(color = country))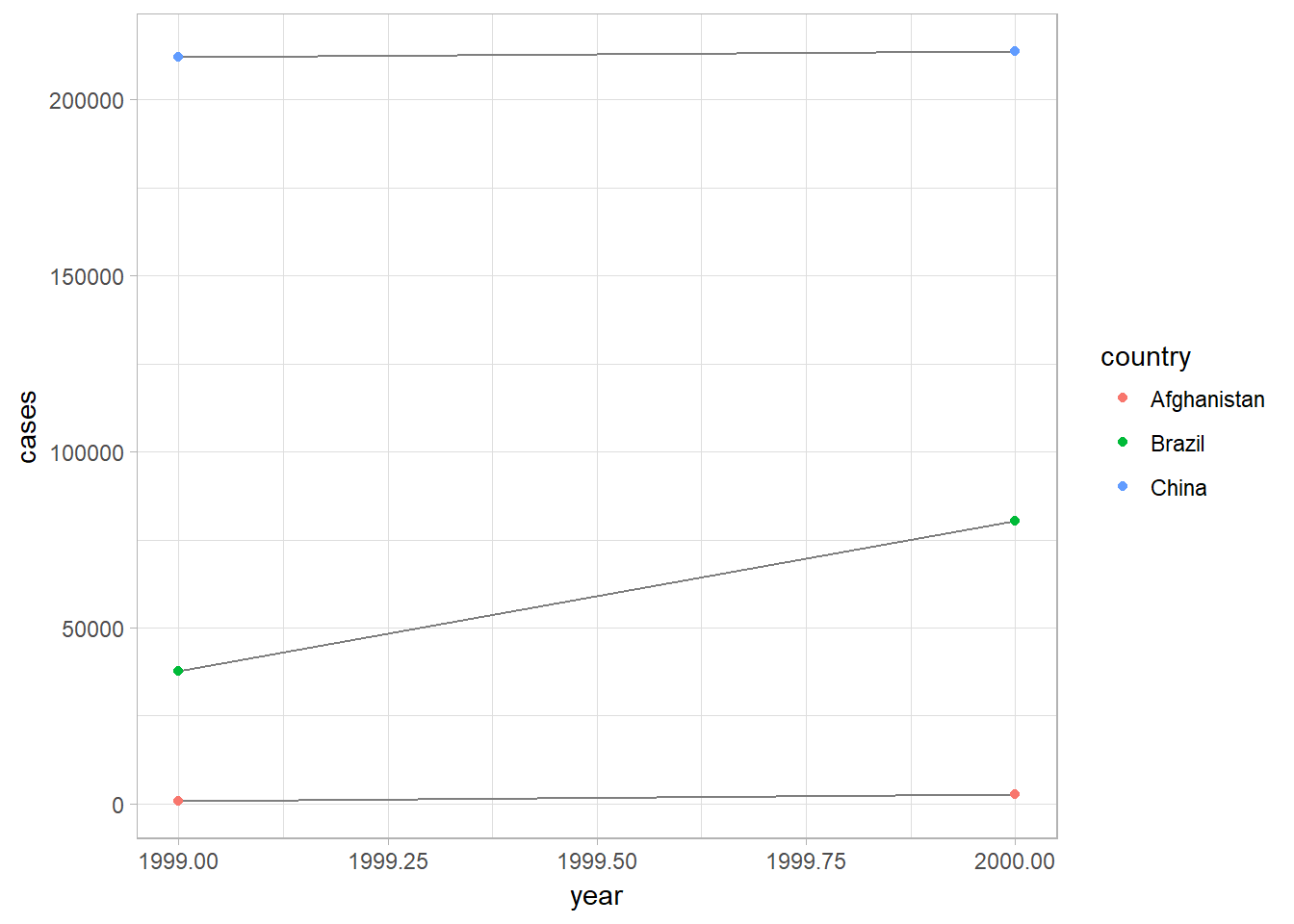
Spreading and Gathering
Most data that you will encounter will be untidy. There are two main reasons:
- Most people aren’t familiar with the principles of tidy data, and it’s hard to derive them yourself unless you spend a lot of time working with data.
- Data is often organized to facilitate some use other than analysis. For example, data is often organized to make entry as easy as possible.
Thus, you need to do some tidying for the most of the datasets:
- Figure out what the variables and observations are.
- Resolve one of two common problems using tidyr’s
gather()andspread():- One variable might be spread across multiple columns. Use
gather()for this. - One observation might be scattered across multiple rows. Use
spread()for this.
- One variable might be spread across multiple columns. Use
Gathering by gather(data, key, value, cols_selection)
A common problem is a dataset where some of the column names are not names of variables, but values of a variable.
For example, column names 1999 and 2000 represent values of the year variable in table4a,and each row represents two observations, not one:
table4a## # A tibble: 3 x 3
## country `1999` `2000`
## * <chr> <int> <int>
## 1 Afghanistan 745 2666
## 2 Brazil 37737 80488
## 3 China 212258 213766To tidy a dataset like this, we need to gather those columns into a new pair of variables.
gather(data, value_column, key_var_name, spreading_value)
- The set of columns that represent values, not variables. In this example, those are the columns
1999and2000. - The name of the variable whose values form the column names. I call that the key, and here it is
year. - The name of the variable whose values are spread over the cells. I call that
value, and here it’s the number ofcases.
table4a %>%
`gather(`1999`, `2000`, key = "year", value = "cases")
table4b %>%
gather(`1999`, `2000`, key = "year", value = "population")## Error: <text>:2:10: unexpected numeric constant
## 1: table4a %>%
## 2: `gather(`1999
## ^To combine the tidied versions of table4a and table4b into a single tibble, we need to use dplyr::left_join():
tidy4a <- table4a %>%
gather(`1999`, `2000`, key = "year", value = "cases")
tidy4b <- table4b %>%
gather(`1999`, `2000`, key = "year", value = "population")
left_join(tidy4a, tidy4b)## # A tibble: 6 x 4
## country year cases population
## <chr> <chr> <int> <int>
## 1 Afghanistan 1999 745 19987071
## 2 Brazil 1999 37737 172006362
## 3 China 1999 212258 1272915272
## 4 Afghanistan 2000 2666 20595360
## 5 Brazil 2000 80488 174504898
## 6 China 2000 213766 1280428583Spreading with spread(data, key, value, fill = NA)
Spreading is the opposite of gathering. You use it when an observation is scattered across multiple rows. Thus, spread and gather are complement.
For example, take table2-an observation is a country in a year, but each observation is spread across two rows:
table2## # A tibble: 12 x 4
## country year type count
## <chr> <int> <chr> <int>
## 1 Afghanistan 1999 cases 745
## 2 Afghanistan 1999 population 19987071
## 3 Afghanistan 2000 cases 2666
## 4 Afghanistan 2000 population 20595360
## 5 Brazil 1999 cases 37737
## 6 Brazil 1999 population 172006362
## 7 Brazil 2000 cases 80488
## 8 Brazil 2000 population 174504898
## 9 China 1999 cases 212258
## 10 China 1999 population 1272915272
## 11 China 2000 cases 213766
## 12 China 2000 population 1280428583The syntax of spread is:
spread(dataframe, key = col_name_or_position_of_var, value = cols_with_multiple_vars_values, fill = NA)
spread(table2, key = type, value = count)## # A tibble: 6 x 4
## country year cases population
## <chr> <int> <int> <int>
## 1 Afghanistan 1999 745 19987071
## 2 Afghanistan 2000 2666 20595360
## 3 Brazil 1999 37737 172006362
## 4 Brazil 2000 80488 174504898
## 5 China 1999 212258 1272915272
## 6 China 2000 213766 1280428583fill = ensures that missing values will be replaced by this value.
spread() and gather() are complements. gather() makes wide tables narrower and longer; spread() makes long tables shorter and wider.
Unique identifiers in spread
Spreading this tibble would fail:
people <- tribble(
~name, ~key, ~value,
#-----------------|--------|------
"Phillip Woods", "age", 45,
"Phillip Woods", "height", 186,
"Phillip Woods", "age", 50,
"Jessica Cordero", "age", 37,
"Jessica Cordero", "height", 156
)
spread(people, key, value)## Error: Duplicate identifiers for rows (1, 3)Spreading the tibble fails because there are two rows with “age” for “Phillip Woods”. We would need to add another column with an indicator for the number observation it is:
people[["id"]] <- c(1,1,2,3,3)
spread(people, key, value)## # A tibble: 3 x 4
## name id age height
## <chr> <dbl> <dbl> <dbl>
## 1 Jessica Cordero 3. 37. 156.
## 2 Phillip Woods 1. 45. 186.
## 3 Phillip Woods 2. 50. NAImperfect symmetrical complement of spread and gather| unless convert = TRUE
For example:
stocks <- tibble(
year = c(2015, 2015, 2016, 2016),
half = c( 1, 2, 1, 2),
return = c(1.88, 0.59, 0.92, 0.17)
)
stocks %>%
spread(year, return) %>%
gather("year", "return", `2015`:`2016`)## # A tibble: 4 x 3
## half year return
## <dbl> <chr> <dbl>
## 1 1. 2015 1.88
## 2 2. 2015 0.590
## 3 1. 2016 0.920
## 4 2. 2016 0.170The functions spread and gather are not perfectly symmetrical because column type information is not transferred between them. In the original table the column year was numeric, but after running spread() and gather() it is a character vector. This is because the key variable names are always converted to a character vector by gather() or it will be saved as factor if the factor_key = TRUE.
The convert argument tries to convert character vectors to the appropriate type. In the background this uses the type.convert function.
stocks %>%
spread(year, return) %>%
gather("year", "return", `2015`:`2016`, convert = TRUE)## # A tibble: 4 x 3
## half year return
## <dbl> <int> <dbl>
## 1 1. 2015 1.88
## 2 2. 2015 0.590
## 3 1. 2016 0.920
## 4 2. 2016 0.170Separating and Uniting
Separate by separate(data, ... = col_to_be_separated, into = new_vars_name)
separate() pulls apart one column into multiple columns, by splitting wherever a separator character appears.
The rate column in table3 contains both cases and population variables and we need to split it into two variables.
separate(data, col_to_be_separated, into = new_vars_name)
table3 %>%
separate(rate, into = c("cases", "population"))## # A tibble: 6 x 4
## country year cases population
## * <chr> <int> <chr> <chr>
## 1 Afghanistan 1999 745 19987071
## 2 Afghanistan 2000 2666 20595360
## 3 Brazil 1999 37737 172006362
## 4 Brazil 2000 80488 174504898
## 5 China 1999 212258 1272915272
## 6 China 2000 213766 1280428583Type conversion by convert = TRUE
Note that the default behavior in separate(): it leaves the type of the column as it originally is. However, We can ask separate() to try and convert to better types using convert = TRUE:
table3 %>%
separate(
rate,
into = c("cases", "population"),
convert = TRUE
)## # A tibble: 6 x 4
## country year cases population
## * <chr> <int> <int> <int>
## 1 Afghanistan 1999 745 19987071
## 2 Afghanistan 2000 2666 20595360
## 3 Brazil 1999 37737 172006362
## 4 Brazil 2000 80488 174504898
## 5 China 1999 212258 1272915272
## 6 China 2000 213766 1280428583Specific character as split values by sep = "/"
By default, separate() will split values wherever it sees a nonalphanumeric character (i.e., a character that isn’t a number or letter). If you wish to use a specific character to separate a column, you can pass the character to the sep argument of separate().
table3 %>%
separate(rate, into = c("cases", "population"), sep = "/")## # A tibble: 6 x 4
## country year cases population
## * <chr> <int> <chr> <chr>
## 1 Afghanistan 1999 745 19987071
## 2 Afghanistan 2000 2666 20595360
## 3 Brazil 1999 37737 172006362
## 4 Brazil 2000 80488 174504898
## 5 China 1999 212258 1272915272
## 6 China 2000 213766 1280428583(Formally, sep is a regular expression)
Split by vector of integers by sep = c(2,4)
You can also pass a vector of integers to sep. separate() will interpret the integers as positions to split at:
- Positive values start at 1 on the far left of the strings;
- negative values start at -1 on the far right of the strings.
When using integers to separate strings, the length of sep should be one less than the number of names in into. For example,
table3 %>%
separate(year, into = c("century", "decade", "year"), sep = c(2,3))## # A tibble: 6 x 5
## country century decade year rate
## * <chr> <chr> <chr> <chr> <chr>
## 1 Afghanistan 19 9 9 745/19987071
## 2 Afghanistan 20 0 0 2666/20595360
## 3 Brazil 19 9 9 37737/172006362
## 4 Brazil 20 0 0 80488/174504898
## 5 China 19 9 9 212258/1272915272
## 6 China 20 0 0 213766/1280428583extra in separate
The extra argument tells separate what to do if there are too many pieces.
For example:
tibble(x = c("a,b,c", "d,e,f,g", "h,i,j"))## # A tibble: 3 x 1
## x
## <chr>
## 1 a,b,c
## 2 d,e,f,g
## 3 h,i,jWe can see that we have an extra g.
By default separate drops the extra values with a warning:
tibble(x = c("a,b,c", "d,e,f,g", "h,i,j")) %>%
separate(x, c("one", "two", "three"))## # A tibble: 3 x 3
## one two three
## <chr> <chr> <chr>
## 1 a b c
## 2 d e f
## 3 h i jThe following produces the same result as above, dropping extra values, but without the warning:
tibble(x = c("a,b,c", "d,e,f,g", "h,i,j")) %>%
separate(x, c("one", "two", "three"), extra = "drop")## # A tibble: 3 x 3
## one two three
## <chr> <chr> <chr>
## 1 a b c
## 2 d e f
## 3 h i jIn the following, the extra values are not split, so “f,g” appears in column three:
tibble(x = c("a,b,c", "d,e,f,g", "h,i,j")) %>%
separate(x, c("one", "two", "three"), extra = "merge")## # A tibble: 3 x 3
## one two three
## <chr> <chr> <chr>
## 1 a b c
## 2 d e f,g
## 3 h i jfill in separate
The fill argument tells separate what to do if there aren’t enough pieces. Now, let’s look at another tibble where we don’t have enough entries (g is missing in this case):
tibble(x = c("a,b,c", "d,e", "f,g,i"))## # A tibble: 3 x 1
## x
## <chr>
## 1 a,b,c
## 2 d,e
## 3 f,g,iThe default for fill is similar to separate; it fills with missing values but emits a warning. In this, row 2 of column “three”, is NA:
tibble(x = c("a,b,c", "d,e", "f,g,i")) %>%
separate(x, c("one", "two", "three"))## # A tibble: 3 x 3
## one two three
## <chr> <chr> <chr>
## 1 a b c
## 2 d e <NA>
## 3 f g iAlternative options for fill are "right", to fill with missing values from the right, but without a warning:
tibble(x = c("a,b,c", "d,e", "f,g,i")) %>%
separate(x, c("one", "two", "three"), fill = "right")## # A tibble: 3 x 3
## one two three
## <chr> <chr> <chr>
## 1 a b c
## 2 d e <NA>
## 3 f g iThe option fill = "left" also fills with missing values without a warning, but this time from the left side. Now, column “one” of row 2 will be missing, and the other values in that row are shifted over:
tibble(x = c("a,b,c", "d,e", "f,g,i")) %>%
separate(x, c("one", "two", "three"), fill = "left")## # A tibble: 3 x 3
## one two three
## <chr> <chr> <chr>
## 1 a b c
## 2 <NA> d e
## 3 f g iUnite with unite(data, col = new_col_name, ... = selection_of_cols, sep = "_", remove = TRUE)
unite() is the inverse of separate(): it combines multiple columns into a single column.
We can use unite() to rejoin the century and year columns in table5.
unite(data, col = new_col_name, ... = selection_of_cols, sep = "_", remove = TRUE)
remove If TRUE, remove input columns from output data frame.
table5 %>%
unite(col = new, century, year, sep = "")## # A tibble: 6 x 3
## country new rate
## <chr> <chr> <chr>
## 1 Afghanistan 1999 745/19987071
## 2 Afghanistan 2000 2666/20595360
## 3 Brazil 1999 37737/172006362
## 4 Brazil 2000 80488/174504898
## 5 China 1999 212258/1272915272
## 6 China 2000 213766/1280428583remove in unite() and separate()
Both unite() and separate() have a remove argument. It will remove the old variables if it is set to TRUE. You would set it to FALSE if you want to create a new variable while keeping the old one.
Missing Values
Surprisingly, a value can be missing in one of two possible ways:
- Explicitly, i.e., flagged with NA.
- Implicitly, i.e., simply not present in the data.
For example:
stocks <- tibble(
year = c(2015, 2015, 2015, 2015, 2016, 2016, 2016),
qtr = c( 1, 2, 3, 4, 2, 3, 4),
return = c(1.88, 0.59, 0.35, NA, 0.92, 0.17, 2.66)
)There are two missing values in this dataset:
- The return for the fourth quarter of 2015 is explicitly missing, because the cell where its value should be instead contains NA.
- The return for the first quarter of 2016 is implicitly missing, because it simply does not appear in the dataset.
The way that a dataset is represented can make implicit values explicit. For example, we can make the implicit missing value explicit by putting years in the columns:
stocks %>%
spread(year, return)## # A tibble: 4 x 3
## qtr `2015` `2016`
## <dbl> <dbl> <dbl>
## 1 1. 1.88 NA
## 2 2. 0.590 0.920
## 3 3. 0.350 0.170
## 4 4. NA 2.66Turning explicit to implicit missing values using na.rm = TRUE
Because these explicit missing values may not be important in other representations of the data, you can set na.rm = TRUE in gather() to turn explicit missing values implicit:
stocks %>%
spread(year, return) %>%
gather(year, return, `2015`:`2016`, na.rm = TRUE)## # A tibble: 6 x 3
## qtr year return
## * <dbl> <chr> <dbl>
## 1 1. 2015 1.88
## 2 2. 2015 0.590
## 3 3. 2015 0.350
## 4 2. 2016 0.920
## 5 3. 2016 0.170
## 6 4. 2016 2.66Filling in explicit NAs with complete()
complete() takes a set of columns, and finds all unique combinations. It then ensures the original dataset contains all those values, filling in explicit NAs where necessary:
stocks %>%
complete(year, qtr)## # A tibble: 8 x 3
## year qtr return
## <dbl> <dbl> <dbl>
## 1 2015. 1. 1.88
## 2 2015. 2. 0.590
## 3 2015. 3. 0.350
## 4 2015. 4. NA
## 5 2016. 1. NA
## 6 2016. 2. 0.920
## 7 2016. 3. 0.170
## 8 2016. 4. 2.66Filling in missing values from recent entries with fill(data, column)
When a data source has primarily been used for data entry, missing values may indicate that the previous value should be carried forward:
treatment <- tribble(
~ person, ~ treatment, ~response,
"Derrick Whitmore", 1, 7,
NA, 2, 10,
NA, 3, 9,
"Katherine Burke", 1, 4
)You can fill in these missing values with fill(). It takes a set of columns where you want missing values to be replaced by the most recent nonmissing value (sometimes called last observation carried forward):
treatment %>%
fill(person)## # A tibble: 4 x 3
## person treatment response
## <chr> <dbl> <dbl>
## 1 Derrick Whitmore 1. 7.
## 2 Derrick Whitmore 2. 10.
## 3 Derrick Whitmore 3. 9.
## 4 Katherine Burke 1. 4.direction in fill()
fill(data, ..., .direction = c("down", "up"))
With fill, it determines whether NA values should be replaced by the previous non-missing value ("down"), which is the default, or the next non-missing value ("up").
Comparing the fill arguments of spread() and complete()
`spread(data, key, value, fill = NA vs complete(data, ..., fill = list())
- Both replace both explicit and implicit NA.
spread(): If the tidy structure creates combinations of variables that do not exist in the original data set, spread() will place an NA in the resulting cells. NA is R’s missing value symbol. You can change this behaviour by passing fill an alternative value to use.complete()A named list that for each variable supplies a single value to use instead of NA for missing combinations.- For example:
df <- tibble( group = c(1:2, 1), item_id = c(1:2, 2), item_name = c("a", "b", "b"), value1 = 1:3, value2 = 4:6 ) ## W/O specifying the fill df %>% complete(group, nesting(item_id, item_name))## # A tibble: 4 x 5 ## group item_id item_name value1 value2 ## <dbl> <dbl> <chr> <int> <int> ## 1 1. 1. a 1 4 ## 2 1. 2. b 3 6 ## 3 2. 1. a NA NA ## 4 2. 2. b 2 5## Specify a list for the missing value in value1 df %>% complete(group, nesting(item_id, item_name), fill = list(value1 = 0))## # A tibble: 4 x 5 ## group item_id item_name value1 value2 ## <dbl> <dbl> <chr> <dbl> <int> ## 1 1. 1. a 1. 4 ## 2 1. 2. b 3. 6 ## 3 2. 1. a 0. NA ## 4 2. 2. b 2. 5
# Case study
The tidyr::who dataset contains tuberculosis (TB) cases broken down by year, country, age, gender, and diagnosis method. Let’ take a look at the dataset:
who## # A tibble: 7,240 x 60
## country iso2 iso3 year new_sp_m014 new_sp_m1524 new_sp_m2534
## <chr> <chr> <chr> <int> <int> <int> <int>
## 1 Afghanistan AF AFG 1980 NA NA NA
## 2 Afghanistan AF AFG 1981 NA NA NA
## 3 Afghanistan AF AFG 1982 NA NA NA
## 4 Afghanistan AF AFG 1983 NA NA NA
## 5 Afghanistan AF AFG 1984 NA NA NA
## 6 Afghanistan AF AFG 1985 NA NA NA
## 7 Afghanistan AF AFG 1986 NA NA NA
## 8 Afghanistan AF AFG 1987 NA NA NA
## 9 Afghanistan AF AFG 1988 NA NA NA
## 10 Afghanistan AF AFG 1989 NA NA NA
## # ... with 7,230 more rows, and 53 more variables: new_sp_m3544 <int>,
## # new_sp_m4554 <int>, new_sp_m5564 <int>, new_sp_m65 <int>,
## # new_sp_f014 <int>, new_sp_f1524 <int>, new_sp_f2534 <int>,
## # new_sp_f3544 <int>, new_sp_f4554 <int>, new_sp_f5564 <int>,
## # new_sp_f65 <int>, new_sn_m014 <int>, new_sn_m1524 <int>,
## # new_sn_m2534 <int>, new_sn_m3544 <int>, new_sn_m4554 <int>,
## # new_sn_m5564 <int>, new_sn_m65 <int>, new_sn_f014 <int>,
## # new_sn_f1524 <int>, new_sn_f2534 <int>, new_sn_f3544 <int>,
## # new_sn_f4554 <int>, new_sn_f5564 <int>, new_sn_f65 <int>,
## # new_ep_m014 <int>, new_ep_m1524 <int>, new_ep_m2534 <int>,
## # new_ep_m3544 <int>, new_ep_m4554 <int>, new_ep_m5564 <int>,
## # new_ep_m65 <int>, new_ep_f014 <int>, new_ep_f1524 <int>,
## # new_ep_f2534 <int>, new_ep_f3544 <int>, new_ep_f4554 <int>,
## # new_ep_f5564 <int>, new_ep_f65 <int>, newrel_m014 <int>,
## # newrel_m1524 <int>, newrel_m2534 <int>, newrel_m3544 <int>,
## # newrel_m4554 <int>, newrel_m5564 <int>, newrel_m65 <int>,
## # newrel_f014 <int>, newrel_f1524 <int>, newrel_f2534 <int>,
## # newrel_f3544 <int>, newrel_f4554 <int>, newrel_f5564 <int>,
## # newrel_f65 <int>It contains redundant columns odd variable codes, and many missing values. In short, who is messy, and we’ll need multiple steps to tidy it.
The best place to start is almost always to gather together the columns that are not variables. Let’s have a look at what we’ve got:
- It looks like
country,iso2, andiso3are three variables that redundantly specify the country. - year is clearly also a variable.
- We don’t know what all the other columns are yet, but given the structure in the variable names (e.g.,
new_sp_m014,new_ep_m014,new_ep_f014) these are likely to be values, not variables.
So we need to gather together all the columns from new_sp_m014 to newrel_f65. We don’t know what those values represent yet, so we’ll give them the generic name "key". We know the cells represent the count of cases, so we’ll use the variable cases. There are a lot of missing values in the current representation, so for now we’ll use na.rm just so we can focus on the values that are present:
who1 <- who %>%
gather(new_sp_m014:newrel_f65,
key = "key", value = "cases", na.rm = TRUE)
who1## # A tibble: 76,046 x 6
## country iso2 iso3 year key cases
## * <chr> <chr> <chr> <int> <chr> <int>
## 1 Afghanistan AF AFG 1997 new_sp_m014 0
## 2 Afghanistan AF AFG 1998 new_sp_m014 30
## 3 Afghanistan AF AFG 1999 new_sp_m014 8
## 4 Afghanistan AF AFG 2000 new_sp_m014 52
## 5 Afghanistan AF AFG 2001 new_sp_m014 129
## 6 Afghanistan AF AFG 2002 new_sp_m014 90
## 7 Afghanistan AF AFG 2003 new_sp_m014 127
## 8 Afghanistan AF AFG 2004 new_sp_m014 139
## 9 Afghanistan AF AFG 2005 new_sp_m014 151
## 10 Afghanistan AF AFG 2006 new_sp_m014 193
## # ... with 76,036 more rowsWe can get some hint of the structure of the values in the new key column by counting them:
who1 %>%
count(key)## # A tibble: 56 x 2
## key n
## <chr> <int>
## 1 new_ep_f014 1032
## 2 new_ep_f1524 1021
## 3 new_ep_f2534 1021
## 4 new_ep_f3544 1021
## 5 new_ep_f4554 1017
## 6 new_ep_f5564 1017
## 7 new_ep_f65 1014
## 8 new_ep_m014 1038
## 9 new_ep_m1524 1026
## 10 new_ep_m2534 1020
## # ... with 46 more rowsThe data dictionary tells us: 1. The first three letters of each column denote whether the column contains new or old cases of TB. In this dataset, each column contains new cases. 2. The next two letters describe the type of TB: i) rel stands for cases of relapse. ii) ep stands for cases of extrapulmonary TB. iii) sn stands for cases of pulmonary TB that could not be diagnosed by a pulmonary smear (smear negative). iv) sp stands for cases of pulmonary TB that could be diagnosed be a pulmonary smear (smear positive). 3. The sixth letter gives the sex of TB patients. The dataset groups cases by males (m) and females (f). 4. The remaining numbers give the age group. The dataset groups cases into seven age groups: i) 014 = 0-14 years old ii) 1524 = 15-24 years old iii) 2534 = 25-34 years old iv) 3544 = 35-44 years old v) 4554 = 45-54 years old vi) 5564 = 55-64 years old vii) 65 = 65 or older
We are fixing a minor mistake here: inconsistent column name between newrel and new_rel.
who2 <- who1 %>%
mutate(key = stringr::str_replace(key, "newrel", "new_rel"))
who2## # A tibble: 76,046 x 6
## country iso2 iso3 year key cases
## <chr> <chr> <chr> <int> <chr> <int>
## 1 Afghanistan AF AFG 1997 new_sp_m014 0
## 2 Afghanistan AF AFG 1998 new_sp_m014 30
## 3 Afghanistan AF AFG 1999 new_sp_m014 8
## 4 Afghanistan AF AFG 2000 new_sp_m014 52
## 5 Afghanistan AF AFG 2001 new_sp_m014 129
## 6 Afghanistan AF AFG 2002 new_sp_m014 90
## 7 Afghanistan AF AFG 2003 new_sp_m014 127
## 8 Afghanistan AF AFG 2004 new_sp_m014 139
## 9 Afghanistan AF AFG 2005 new_sp_m014 151
## 10 Afghanistan AF AFG 2006 new_sp_m014 193
## # ... with 76,036 more rowsThen, we separate the key into several columns by seperator _ :
who3 <- who2 %>%
separate(key, c("new", "type", "sexage"), sep = "_")
who3## # A tibble: 76,046 x 8
## country iso2 iso3 year new type sexage cases
## <chr> <chr> <chr> <int> <chr> <chr> <chr> <int>
## 1 Afghanistan AF AFG 1997 new sp m014 0
## 2 Afghanistan AF AFG 1998 new sp m014 30
## 3 Afghanistan AF AFG 1999 new sp m014 8
## 4 Afghanistan AF AFG 2000 new sp m014 52
## 5 Afghanistan AF AFG 2001 new sp m014 129
## 6 Afghanistan AF AFG 2002 new sp m014 90
## 7 Afghanistan AF AFG 2003 new sp m014 127
## 8 Afghanistan AF AFG 2004 new sp m014 139
## 9 Afghanistan AF AFG 2005 new sp m014 151
## 10 Afghanistan AF AFG 2006 new sp m014 193
## # ... with 76,036 more rowsIt’s found at below that the new column is redundant because they are all the same:
who3 %>%
count(new)## # A tibble: 1 x 2
## new n
## <chr> <int>
## 1 new 76046In addition, iso2 and iso3 is redundant because it is tied to country name. We can show that by counting the occurences of unique combination of country, iso2, and iso3; if the occurences is not more than one, then iso are uniwue for each country:
who3 %>%
select(country, iso2, iso3) %>%
distinct() %>%
group_by(country) %>%
filter(n()>1)## # A tibble: 0 x 3
## # Groups: country [0]
## # ... with 3 variables: country <chr>, iso2 <chr>, iso3 <chr>Let’s drop the new column alongside with iso2 and iso3 since they’re redundant:
who4 <- who3 %>%
select(-new, - iso2, -iso3)Then, we separate sex and age from sexage by splitting after the first character:
who5 <- who4%>%
separate(sexage, into = c("sex", "age"), sep = 1)
who5## # A tibble: 76,046 x 6
## country year type sex age cases
## <chr> <int> <chr> <chr> <chr> <int>
## 1 Afghanistan 1997 sp m 014 0
## 2 Afghanistan 1998 sp m 014 30
## 3 Afghanistan 1999 sp m 014 8
## 4 Afghanistan 2000 sp m 014 52
## 5 Afghanistan 2001 sp m 014 129
## 6 Afghanistan 2002 sp m 014 90
## 7 Afghanistan 2003 sp m 014 127
## 8 Afghanistan 2004 sp m 014 139
## 9 Afghanistan 2005 sp m 014 151
## 10 Afghanistan 2006 sp m 014 193
## # ... with 76,036 more rowsThe dataset is tidy now!
Nontidy Data
There are lots of useful and well-founded data structures that are not tidy data. There are two main reasons to use other data structures:
- Alternative representations may have substantial performance or space advantages.
- Specialized fields have evolved their own conventions for storing data that may be quite different to the conventions of tidy data.
Tidy data should be your choice if your data fits into rectangular structure.